Ribosome Biogenesis on:
[Wikipedia]
[Google]
[Amazon]
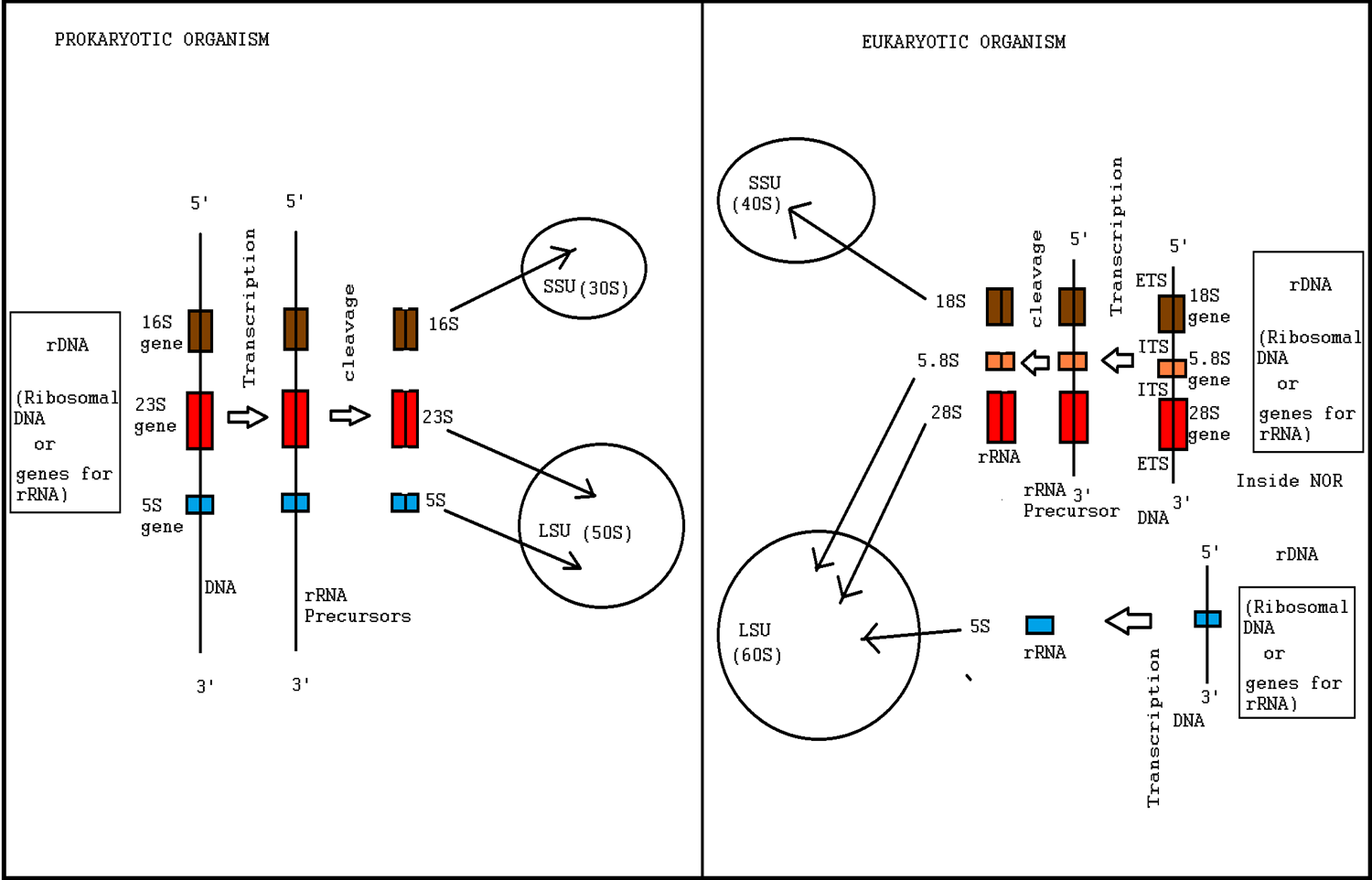 Ribosome biogenesis is the process of making ribosomes. In
Ribosome biogenesis is the process of making ribosomes. In
''GTP + ATP --> pppGpp + AMP'' The γ-phosphate is then removed and ppGpp will bind to and inhibit RNA polymerase. This binding causes a reduction in rRNA transcription. A reduced amount of rRNA means that ribosomal proteins (r-proteins) will be translated but will not have an rRNA to bind to. Instead, they will negatively feedback and bind to their ''own'' mRNA, repressing r-protein synthesis. Note that r-proteins preferentially bind to their complementary rRNA if it is present, rather than mRNA. The ribosome operons also include the genes for RNA polymerase and
 Ribosome biogenesis is the process of making ribosomes. In
Ribosome biogenesis is the process of making ribosomes. In prokaryote
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Conne ...
s, this process takes place in the cytoplasm with the transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
of many ribosome gene operon
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splic ...
s. In eukaryotes, it takes place both in the cytoplasm
In cell biology, the cytoplasm is all of the material within a eukaryotic cell, enclosed by the cell membrane, except for the cell nucleus. The material inside the nucleus and contained within the nuclear membrane is termed the nucleoplasm. ...
and in the nucleolus
The nucleolus (, plural: nucleoli ) is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis, which is the synthesis of ribosomes. The nucleolus also participates in the formation of ...
. It involves the coordinated function of over 200 proteins in the synthesis and processing of the three prokaryotic
A prokaryote () is a single-celled organism that lacks a nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Greek πρό (, 'before') and κάρυον (, 'nut' or 'kernel').Campbell, N. "Biology:Concepts & Connec ...
or four eukaryotic
Eukaryotes () are organisms whose Cell (biology), cells have a cell nucleus, nucleus. All animals, plants, fungi, and many unicellular organisms, are Eukaryotes. They belong to the group of organisms Eukaryota or Eukarya, which is one of the ...
rRNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosoma ...
s, as well as assembly of those rRNAs with the ribosomal proteins. Most of the ribosomal proteins fall into various energy-consuming enzyme families including ATP-dependent RNA helicase
Helicases are a class of enzymes thought to be vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separatin ...
s, AAA-ATPases, GTPase
GTPases are a large family of hydrolase enzymes that bind to the nucleotide guanosine triphosphate (GTP) and hydrolyze it to guanosine diphosphate (GDP). The GTP binding and hydrolysis takes place in the highly conserved P-loop "G domain", a pro ...
s, and kinases
In biochemistry, a kinase () is an enzyme that catalyzes the transfer of phosphate groups from high-energy, phosphate-donating molecules to specific substrates. This process is known as phosphorylation, where the high-energy ATP molecule don ...
. About 60% of a cell's energy is spent on ribosome production and maintenance.
Ribosome biogenesis is a very tightly regulated process, and it is closely linked to other cellular activities like growth and division.
Some have speculated that in the origin of life, ribosome biogenesis predates cells, and that genes and cells evolved to enhance the reproductive capacity of ribosomes.
Ribosomes
Ribosomes
Ribosomes ( ) are macromolecular machines, found within all cells, that perform biological protein synthesis (mRNA translation). Ribosomes link amino acids together in the order specified by the codons of messenger RNA (mRNA) molecules to ...
are the macromolecular machines that are responsible for mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
translation
Translation is the communication of the meaning of a source-language text by means of an equivalent target-language text. The English language draws a terminological distinction (which does not exist in every language) between ''transla ...
into proteins. The eukaryotic ribosome, also called the 80S ribosome, is made up of two subunits – the large 60S subunit (which contains the 25S n plantsor 28S n mammals 5.8S, and 5S rRNA and 46 ribosomal proteins) and a small 40S subunit (which contains the 18S rRNA and 33 ribosomal proteins). The ribosomal proteins are encoded by ribosomal genes.
Prokaryotes
There are 52 genes that encode the ribosomal proteins, and they can be found in 20operons
In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splic ...
within prokaryotic DNA. Regulation of ribosome synthesis hinges on the regulation of the rRNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosoma ...
itself.
First, a reduction in aminoacyl-tRNA
Aminoacyl-tRNA (also aa-tRNA or charged tRNA) is tRNA to which its cognate amino acid is chemically bonded (charged). The aa-tRNA, along with particular elongation factors, deliver the amino acid to the ribosome for incorporation into the polypept ...
will cause the prokaryotic cell to respond by lowering transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
and translation
Translation is the communication of the meaning of a source-language text by means of an equivalent target-language text. The English language draws a terminological distinction (which does not exist in every language) between ''transla ...
. This occurs through a series of steps, beginning with stringent factors binding to ribosomes and catalyzing the reaction:''GTP + ATP --> pppGpp + AMP'' The γ-phosphate is then removed and ppGpp will bind to and inhibit RNA polymerase. This binding causes a reduction in rRNA transcription. A reduced amount of rRNA means that ribosomal proteins (r-proteins) will be translated but will not have an rRNA to bind to. Instead, they will negatively feedback and bind to their ''own'' mRNA, repressing r-protein synthesis. Note that r-proteins preferentially bind to their complementary rRNA if it is present, rather than mRNA. The ribosome operons also include the genes for RNA polymerase and
elongation factors
Elongation factors are a set of proteins that function at the ribosome, during protein synthesis, to facilitate translational elongation from the formation of the first to the last peptide bond of a growing polypeptide. Most common elongat ...
(used in RNA translation). Regulation of all of these genes at once illustrate the coupling between transcription and translation in prokaryotes.
Eukaryotes
Ribosomal protein synthesis in eukaryotes is a major metabolic activity. It occurs, like most protein synthesis, in the cytoplasm just outside the nucleus. Individual ribosomal proteins are synthesized and imported into the nucleus throughnuclear pores
A nuclear pore is a part of a large complex of proteins, known as a nuclear pore complex that spans the nuclear envelope, which is the double membrane surrounding the eukaryotic cell nucleus. There are approximately 1,000 nuclear pore complexes ...
. See nuclear import A nuclear localization signal ''or'' sequence (NLS) is an amino acid sequence that 'tags' a protein for import into the cell nucleus by nuclear transport. Typically, this signal consists of one or more short sequences of positively charged lysines ...
for more about the movement of the ribosomal proteins into the nucleus.
The DNA is transcribed, at a high speed, in the nucleolus
The nucleolus (, plural: nucleoli ) is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis, which is the synthesis of ribosomes. The nucleolus also participates in the formation of ...
, which contains all 45S rRNA genes. The only exception is the 5S rRNA which is transcribed outside the nucleolus. After transcription, the rRNAs associate with the ribosomal proteins, forming the two types of ribosomal subunits (large and small). These will later assemble in the cytosol to make a functioning ribosome. See nuclear export
A nuclear export signal (NES) is a short target peptide containing 4 hydrophobic residues in a protein that targets it for export from the cell nucleus to the cytoplasm through the nuclear pore complex using nuclear transport. It has the opposite ...
for more about the movement of the ribosomal subunits out of the nucleus.
Processing
Eukaryotic cells co-transcribe three of the mature rRNA species through a series of steps. The maturation process of the rRNAs and the process of recruiting the r-proteins happen in precursor ribosomal particles, sometimes called pre-ribosomes, and takes place in thenucleolus
The nucleolus (, plural: nucleoli ) is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis, which is the synthesis of ribosomes. The nucleolus also participates in the formation of ...
, nucleoplasm
The nucleoplasm, also known as karyoplasm, is the type of protoplasm that makes up the cell nucleus, the most prominent organelle of the eukaryotic cell. It is enclosed by the nuclear envelope, also known as the nuclear membrane. The nucleoplasm ...
, and cytoplasm
In cell biology, the cytoplasm is all of the material within a eukaryotic cell, enclosed by the cell membrane, except for the cell nucleus. The material inside the nucleus and contained within the nuclear membrane is termed the nucleoplasm. ...
. The yeast, ''S. cerevisiae'' is the eukaryotic model organism for the study of ribosome biogenesis.
Ribosome biogenesis starts in the nucleolus. There, the 18S, 5.8S, and 25S subunits of the rRNA are cotranscribed from ribosomal genes as a polycistronic
A cistron is an alternative term for "gene". The word cistron is used to emphasize that genes exhibit a specific behavior in a cis-trans test; distinct positions (or loci) within a genome are cistronic.
History
The words ''cistron'' and ''gene ...
transcript by RNA polymerase I
RNA polymerase 1 (also known as Pol I) is, in higher eukaryotes, the polymerase that only transcribes ribosomal RNA (but not 5S rRNA, which is synthesized by RNA polymerase III), a type of RNA that accounts for over 50% of the total RNA synthesize ...
, and is called 35S pre-RNA.
Transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, the fir ...
of polymerase I starts with a Pol I initiation complex that binds to the rDNA promoter. The formation of this complex requires the help of an upstream activating factor or UAF that associates with TATA-box binding protein and the core factor (CF). Together the two transcription factors allow the RNA pol I complex to bind with the polymerase I initiation factor, Rrn3. As the pol I transcript is produced, approximately 75 small nucleolar ribonucleoparticles (snoRNPs) facilitate the co-transcriptional covalent
A covalent bond is a chemical bond that involves the sharing of electrons to form electron pairs between atoms. These electron pairs are known as shared pairs or bonding pairs. The stable balance of attractive and repulsive forces between atoms ...
modifications of >100 rRNA residues. These snoRNPs control 2’-O-ribose methylation of nucleotides and also assist in the creation of pseudouridine
Pseudouridine (abbreviated by the Greek letter psi- Ψ) is an isomer of the nucleoside uridine in which the uracil is attached via a carbon-carbon instead of a nitrogen-carbon glycosidic bond. (In this configuration, uracil is sometimes referred ...
s. At the 5’ end of rRNA transcripts, small subunit ribosomal proteins (Rps) and non-ribosomal factors assemble with the pre-RNA transcripts to create ball-like knobs. These knobs are the first pre-ribosomal particles in the small (40S) ribosomal subunit pathway. The rRNA transcript is cleaved at the A2 site, and this separates the early 40S pre-ribosome from the remaining pre-rRNA that will combine with large subunit ribosomal proteins (Rpl) and other non-ribosomal factors to create the pre-60S ribosomal particles.
40S subunit
The transcriptional assembly of the 40 S subunit precursor, sometimes referred to as the small subunit processome (SSU) or 90S particle happens in a hierarchical fashion – essentially a stepwise incorporation of the UTP-A, UTP-B, and UTP-C subcomplexes. These subcomplexes are made up of over 30 non ribosomal protein factors, the U3 snoRNP particle, a few Rps proteins, and the 35S pre-rRNA. Their exact role, though has not been discovered. The composition of the pre-40S particle changes drastically once cleavage at the U3 snoRNPA dependent sites (sites A0, A1, and A2) are made. This cleavage event creates the 20S pre-rRNA and causes ribosomal factors to dissociate from the pre-40S particle. U3 is displaced from the nascent 40S by the helicase Dhr1. At this point in the ribosome biogenesis process, the 40S pre-ribosome already shows the “head” and “body” structures of the mature 40S subunit. The 40S pre-ribosome is transported out of the nucleolus and into the cytoplasm. The cytoplasmic 40S pre-ribosome now contains ribosomal proteins, the 20s rRNA and a few non-ribosomal factors. The final formation of the 40S subunit “beak” structure occurs after a phosphorylation anddephosphorylation
In biochemistry, dephosphorylation is the removal of a phosphate (PO43−) group from an organic compound by hydrolysis. It is a reversible post-translational modification. Dephosphorylation and its counterpart, phosphorylation, activate and de ...
event involving the Enp1-Ltv1-Rps3 complex and the kinase, Hrr25. Cleavage of the 20S pre-rRNA at the D-site creates the mature 18s rRNA. This cleavage event is dependent on several non-ribosomal factors such as Nob1, Rio1, Rio2, Tsr1 and Fap7.
60S subunit
The maturation of the pre-60S subunit into a mature 60S subunit requires many biogenesis factors that associate and disassociate. In addition, some assembly factors associate with the 60S subunit while others only interact with it transiently. As an overall trend, the maturation of the pre-60S subunit is marked a gradual decrease in complexity. The subunit matures as it moves from the nucleolus to the cytoplasm and gradually the number oftrans-acting In the field of molecular biology, ''trans''-acting (''trans''-regulatory, ''trans''-regulation), in general, means "acting from a different molecule" (''i.e.'', intermolecular). It may be considered the opposite of ''cis''-acting (''cis''-regula ...
factors are reduced. The maturation of the 60S subunit requires the help of about 80 factors. Eight of these factors are directly involved with the processing of the 27S A3 pre-rRNA, which actually completes the formation of the mature 5’end of the 5.8S rRNA. The A3 factors bind to distant sites on the pre-RNA as well as to each other. Subsequently, they bring areas of rRNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosoma ...
close together and promote the processing of pre-rRNA and the recruitment of ribosomal proteins. Three AAA-type ATPases work to strip the factors from the maturing 60S pre-ribosome. One of the ATPases is a dynein-like Rea1 protein made up of 6 different ATPase domains that form a ring structure. The ring structure is attached to a flexible tail that happens to have a MIDAS (Metal ion-dependent adhesion site) tip. The Rea1 interacts with the 60S pre-ribosome via its ring while two substrates, Ytm1 and Rsa1, interact with Rea1 through its MIDAS tip. The role of these substrates has not yet been defined. Both though, along with their interactions, are removed in the maturation process of the 60S pre-ribosome. The other two ATPases, Rix7 and Drg1 also function to remove assembly factors from the maturing 60S subunit. Helicase
Helicases are a class of enzymes thought to be vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separatin ...
s and GTPases are also involved in the removal of assembly factors and the rearrangement of RNA to form the completed 60S subunit. Once in the cytoplasm (see nuclear export), the 60S subunit further undergoes processing in order to be functional. The rest of the large subunit ribosomal particles associate with the 60S unit and the remaining non-ribosomal assembly factors disassociate. The release of the biogenesis factors is mediated mostly by GTPases such as Lsg1 and ATPases such as Drg1. The precise sequence of these events remains unclear. The pathway of 60S cytoplasmic maturation remains incomplete as far as current knowledge is concerned.
Nuclear export
In order for the pre-ribosomal units to fully mature, they must be exported to thecytoplasm
In cell biology, the cytoplasm is all of the material within a eukaryotic cell, enclosed by the cell membrane, except for the cell nucleus. The material inside the nucleus and contained within the nuclear membrane is termed the nucleoplasm. ...
. To effectively move from the nucleolus to the cytoplasm, the pre-ribosomes interact with export receptors to move through the hydrophobic central channel of the nuclear pore complex. The karyopherin
Karyopherins are proteins involved in transporting molecules between the cytoplasm and the nucleus of a eukaryotic cell. The inside of the nucleus is called the karyoplasm (or nucleoplasm). Generally, karyopherin-mediated transport occurs through ...
Crm1
Exportin 1 (XPO1), also known as chromosomal region maintenance 1 (CRM1), is a eukaryotic protein that mediates the nuclear export of various proteins and RNAs.
History
XPO1 (CRM1) originally was identified in the fission yeast ''Schizosaccharom ...
is the receptor for both ribosomal subunits and mediates export in a Ran-GTP dependent fashion. It recognizes molecules that have leucine
Leucine (symbol Leu or L) is an essential amino acid that is used in the biosynthesis of proteins. Leucine is an α-amino acid, meaning it contains an α- amino group (which is in the protonated −NH3+ form under biological conditions), an α- ...
-rich nuclear export signals. The Crm1 is pulled to the large 60S subunit by the help of an adapter protein called Nmd3. The adapter protein for the 40S unit is unknown. In addition to Crm1, other factors play a role in nuclear export of pre-ribosomes. A general mRNA export receptor, called Mex67, as well as a HEAT-repeating-containing protein, Rrp12, facilitate the export of both subunits. These factors are non-essential proteins and help to optimize the export of the pre-ribosomes since they are large molecules.
Quality control
Becauseribosomes
Ribosomes ( ) are macromolecular machines, found within all cells, that perform biological protein synthesis (mRNA translation). Ribosomes link amino acids together in the order specified by the codons of messenger RNA (mRNA) molecules to ...
are so complex, a certain number of ribosomes are assembled incorrectly and could potentially waste cellular energy and resources when synthesizing non-functional proteins. To prevent this, cells have an active surveillance system to recognize damaged or defective ribosomes and target them for degradation. The surveillance mechanism is in place to detect nonfunctional pre-ribosomes as well as nonfunctional mature ribosomes. In addition, the surveillance system brings the necessary degradation equipment and actually degrades the nonfunctional ribosomes. Pre-ribosomes that build up in the nucleus are destroyed by the exosome, which is a multisubunit complex with exonuclease activity. If defective ribosomal subunits do happen to make it out of the nucleolus and into the cytoplasm, there is a second surveillance system in place there to target malfunctioning ribosomes in the cytoplasm for degradation. Certain mutations in residues of the large ribosome subunit will actually result in RNA decay and thus degradation of the unit. Because the amount of defects that are possible in ribosome assembly are so extensive, it is still unknown as to how the surveillance system detects all defects, but it has been postulated that instead of targeting specific defects, the surveillance system recognizes the consequences of those defects – such as assembly delays. Meaning, if there is a disruption in the assembly or maturation of a mature ribosome, the surveillance system will act as if the subunit is defective.
Human disease
Mutations in ribosome biogenesis are linked to several human ribosomopathygenetic diseases
A genetic disorder is a health problem caused by one or more abnormalities in the genome. It can be caused by a mutation in a single gene (monogenic) or multiple genes (polygenic) or by a chromosomal abnormality. Although polygenic disorders ...
, including inherited bone marrow failure syndromes, which are characterized by a predisposition to cancer
Cancer is a group of diseases involving abnormal cell growth with the potential to invade or spread to other parts of the body. These contrast with benign tumors, which do not spread. Possible signs and symptoms include a lump, abnormal b ...
and a reduced number of blood cells. Ribosomal dysregulation may also play a role in muscle wasting
Muscle atrophy is the loss of skeletal muscle mass. It can be caused by immobility, aging, malnutrition, medications, or a wide range of injuries or diseases that impact the musculoskeletal or nervous system. Muscle atrophy leads to muscle weakness ...
.
See also
* RNA polymeraseReferences